How Do We Learn Dec 2025 Aliveline
What is an “Aliveline”?
Early December 2025 I was feeling a bit lost. Or more specifically, I was feeling low on momentum. I recalled enjoying reading about month-to-master projects where someone takes on a seemingly insane challenge, like trying to become a chess grandmaster in a month. They often don’t succeed, but do sometimes fail in an interesting way as constraints can breed creativity. I decided to do one of these in December 2025, and chose the term “aliveline” in lieu of deadline – the point of these constraints is to excite you about a crazy sounding goal and ultimately to make you feel more alive.
I also believe that Ambitious goals are useful as they illuminate a path that maximizes learning and enjoyment, not as ends in themselves, and starting an aliveline was a way to directly test that belief.
I had zero neuroscience background, but it was time to jump start progress on one of my two main quest(ion)s – understanding how we learn.
So for December I chose to try to have and publish an original insight. I began the aliveline on December 2 and was intending to give myself until January 2, but decided to take a holiday break in the middle & wrapped it up on January 31. I actively worked on this ~3-4h a day (I have a baby at home), so I spent ~100h on this in total.
A more detailed definition of my goal, from the first day:
What I mean by “have and publish an original insight”:
- I have been reading here and there about intelligence – how we learn, how the brain works, how ML works, etc
- I want to publish something that is original relative to the current research AND more importantly something that I would consider novel/interesting today
Did I succeed?
Not really.
I ended up working on the neural dynamics of C. elegans — tiny worms with ~300 neurons, one of the simplest animals with a nervous system. The specific experiment was extending a method to extract low-D neural loops representing locomotion in their brains to freely moving worms (the original paper had found clean loops in immobilized worms). I was able to recover loops in the worm neural traces I tested, though the quality of the loops is questionable as they were significantly noisier. Further tests would be needed to fully explore how loop-like the structures that I found are. The research literature qualitatively agrees – others have also found it difficult to find clean neural dynamics loops in freely moving worms, because there is just so much going on in their brains.
Take a look here if you are interested in the details of the project.
My result confirmed what the field already suspected — not nothing for a first attempt, but not exactly an original insight. And it wasn’t a particularly insightful question either by my standards. It is certainly something that I would have considered novel/interesting at the beginning of December, though.
But I feel kinda successful?
Having an absurd objective like this was extremely motivating. It forced me to take a research-first/question-first approach to learning things, which was way more fun and got me ramped up in the field very quickly. LLMs helped massively on that front, along with the coding front. Perhaps unsurprisingly, I spent most of my time (~75%) just getting to a hypothesis grounded in the literature that I was excited to test:
Genes work at the neuron level, but need to encode behavior (eg locomotion) as a top-down computational constraint on the network. They accomplish this via modifying neurons’ nonlinear feedback control algorithms. This results in certain structures (eg 2D loops for locomotion in C Elegans) appearing with very high probability, despite each overall brain’s neural coordinate system being unique.
And while the final experiment didn’t end up with unambiguously positive results and I had a lot of hiccups along the way, I still feel proud that I was able to go from vague ideas to a real hypothesis & run a real experiment with real results to test one small part of it in just two months. And more importantly, it was fun! I am excited to continue my quest of understanding the brain and also extremely interested in using LLMs to accelerate science more broadly.
That said…
There is much room for improvement
After I finished, I wrote out Reflections on the experience and let it sit for a week or so. Then I had Claude go through my daily working logs, the various repos, all my ChatGPT chats & Codex sessions, and trace my thinking process/workflow and suggest areas for improvement. The major ones were:
Execution outran understanding: I frequently jumped to writing code and testing on datasets before making sure the scientific question was well-formulated. LLMs sometimes made execution a little too easy. Similarly, I felt that my newness to the field along with the time constraint made it hard for me to prioritize deep understanding of the fundamentals (nonlinear dynamics, neuronal dynamics, etc). I don’t regret approaching computational neuroscience in this way because it was so fun. But I do think it makes sense to alternate between periods of deeply studying the fundamentals, open time pondering questions, and this kind of sprint.
LLMs were overused for idea generation, underused as adversaries: Pretty self-explanatory & also a result of being new to the field.
Expert contact came far too late: I reached out to authors of papers/methods I was interested in quite late in the process. I regret not following a more person-first approach to learning:
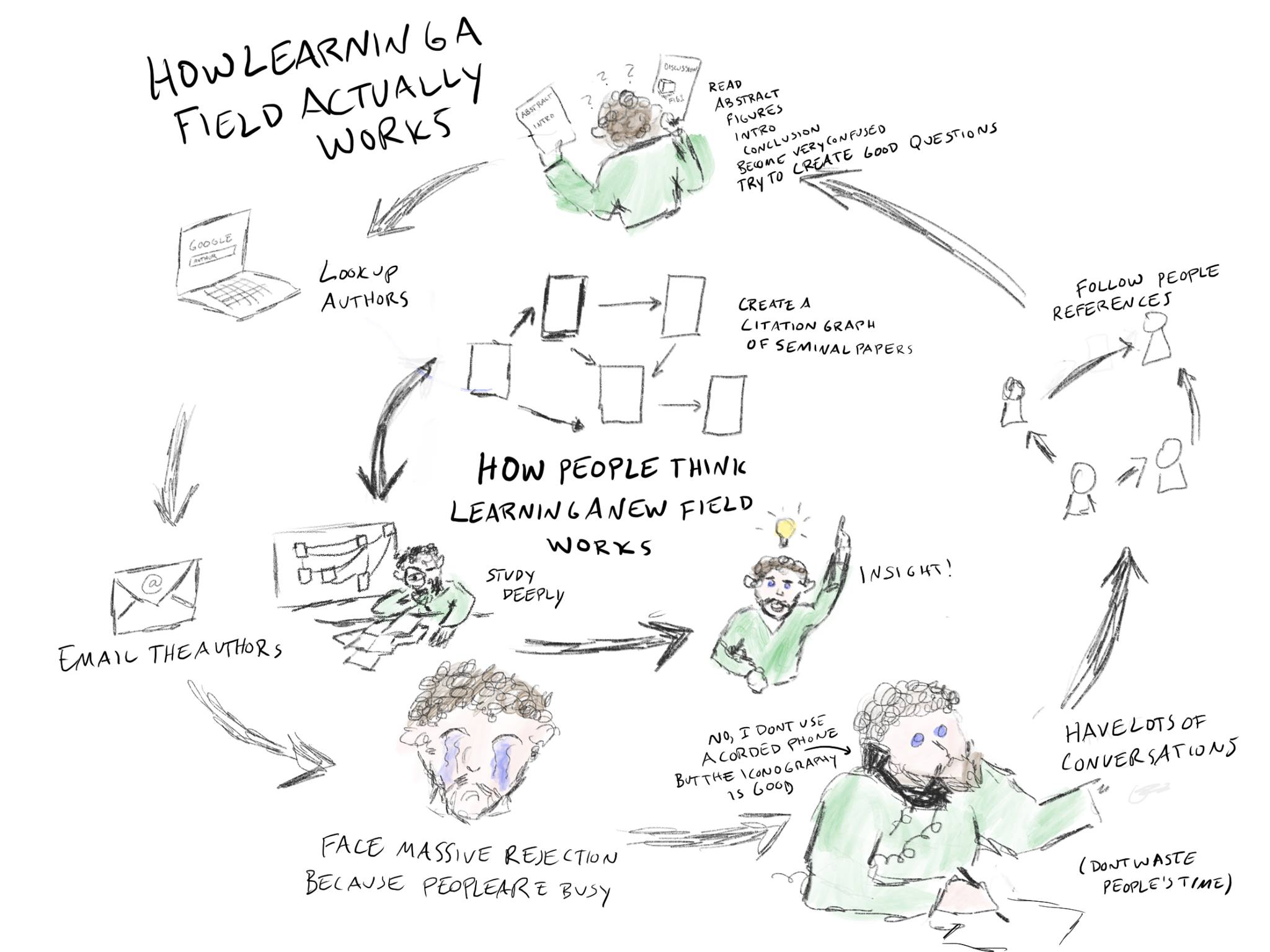
Context fragmented across tools and time: Chat conversations were overly long, Codex sessions were stateless, daily logs weren’t consulted during coding, the 2.5-week holiday break caused significant momentum loss requiring ~1 week to bootstrap. Overall not great management of my own context or the LLM’s.
LLMs could have been less autonomous in certain areas & more so in others: I asked Claude to do a deeper dive on when & how an LLM co-scientist should be more autonomous, based on my chats/daily logs/etc. The mental model of me being the PI, Claude Code being the senior researcher, and Codex being the grad student/builder is one I should have employed.
I am looking forward to implementing these suggestions & also having LLMs help me improve the co-science system itself when I start my next project. I have a feeling that will be quite soon.
If you want to see the struggle
You can get a sense with the arc below, or you can read through my daily working logs – here’s Day 1.
Phase 1: Casting the net (Days 1-6)
- Started with Kolmogorov complexity as a lens on brain function
- Explored Tracy-Widom phase transitions, neural population geometry, the brain as a dynamical system
- Generated many interesting but unfocused ideas
Phase 2: The mouse PFC detour (Days 7-12)
- Settled on the Reinert mouse mPFC dataset as the most tractable test
- Built an eigenvalue analysis pipeline to look for BBP-style phase transitions during rule learning
- Discovered the confound problem is enormous
- Key realization: C. elegans is a better model system for these questions
Phase 3: Deepening into C. elegans (Days 13-20)
- Read the aversive learning paper (Liang et al.), the Kato paper, the Brennan-Proekt paper
- Developed the “macroscopic universality, microscopic degeneracy” framework
- Connected locomotion manifolds, proprioception, and context-gated learning
- Holiday break (Dec 21 - Jan 9)
Phase 4: The DD-DC breakthrough (Days 21-23)
- Read “The neuron as a direct data-driven controller” (Moore et al.)
- Integrated it into a multi-level hypothesis: genes -> feedback control laws -> computational scaffolds -> behavior
- Produced the detailed experiment taxonomy with falsification criteria
Phase 5: Execution and negative results (Days 24-31)
- Applied LOOPER to Kato (positive control) and Atanas (target) datasets
- Discovered: fidelity passes for both, but stationarity fails for both
- Put heat-pulse experiments on hold
- Wrote email with precise questions to LOOPER author